Neuromorphic DevTools
Record. Calibrate.
Sweep. Visualize.
The first experiment tracking platform built for neuromorphic hardware. Import AER events from any chip, characterize transistor mismatch, sweep physical parameters, and visualize everything with one line. Numpy-first. No PyTorch required.
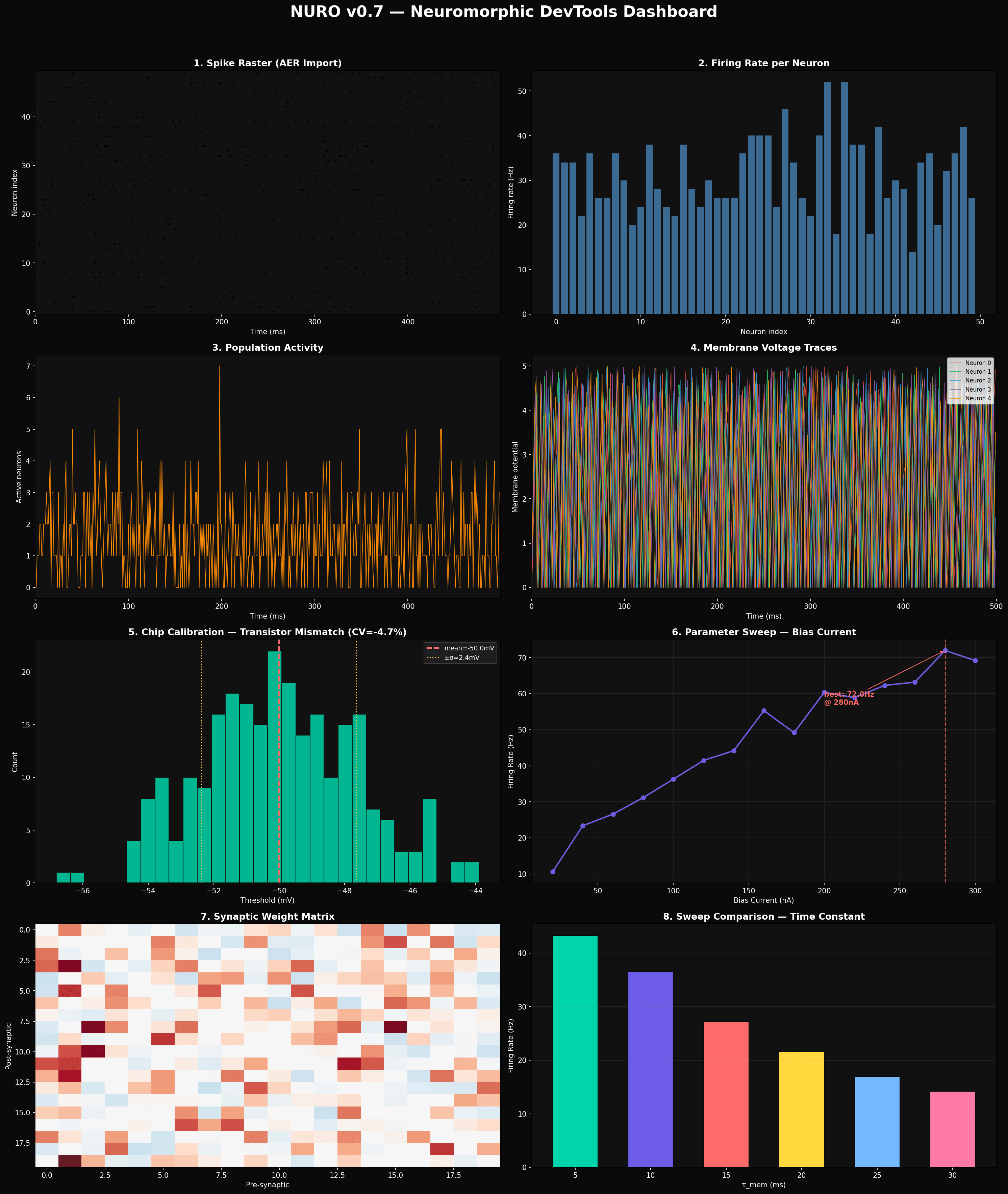
8-panel dashboard generated from real Nuro API calls. Spike raster, firing rates, population activity, voltage traces, chip calibration mismatch, parameter sweep, weight matrix, and sweep comparison.
227
Tests passing
8
Visualization types
6
Hardware adapters
0
PyTorch required
Features
Everything analog and digital neuromorphic labs need for structured experiment tracking. Built from direct researcher feedback.
AER Event Import
Load binary spike data from any neuromorphic chip. Parse aedat 2.0 files from jAER, raw binary AER streams with configurable byte widths, or pre-parsed numpy arrays. Vectorized binning handles 100k+ events instantly.
from nuro.adapters.aer import from_aedat, from_aer_events
# Parse jAER aedat 2.0 files
rec = from_aedat("recording.aedat", num_neurons=256, dt=1e-3)
# From pre-parsed arrays (vectorized)
rec = from_aer_events(neuron_ids, timestamps, num_neurons=256, dt=1e-3)Chip Calibration
Store per-neuron transistor mismatch data for analog neuromorphic chips. Bulk-set parameters across hundreds of neurons, compute mismatch statistics (mean, std, CV, min, max), persist to HDF5. Attach calibration profiles to experiments for full reproducibility.
from nuro.calibration import CalibrationProfile
cal = CalibrationProfile(chip_id="board_03", chip_type="dynap-se2")
cal.set_neurons_bulk("threshold", thresholds) # 256 neurons
cal.set_neurons_bulk("tau", time_constants)
stats = cal.mismatch_stats("threshold")
# {'mean': -50.7, 'std': 2.4, 'cv': -0.049, 'min': -56.2, 'max': -44.1}
exp.set_calibration(cal) # attach to experimentParameter Sweeps
Sweep physical parameters like bias currents, membrane time constants, or firing thresholds. Attach full experiments to each sweep point. Find optimal operating points automatically. Plot sweep curves. Save and load entire sweeps with all associated data.
from nuro.sweep import ParameterSweep
sweep = ParameterSweep(name="bias_sweep", parameter="bias_current", unit="nA")
for bias in [50, 100, 150, 200, 250, 300]:
exp = nuro.experiment(f"run_{bias}")
exp.log_metric("firing_rate_hz", measured_rate)
sweep.add_run(bias, experiment=exp)
best = sweep.best("firing_rate_hz", mode="max")
sweep.plot("firing_rate_hz") # one-line plotExperiment Tracking
Bundle hardware config, calibration profile, recordings, parameters, and metrics into one Experiment object. Save to disk as JSON metadata + HDF5 recordings. Load back with full fidelity. Multi-chip support with per-chip configuration.
exp = nuro.experiment("wta_circuit_v3")
exp.set_hardware("dynap-se2", chips=[
{"chip_id": "board_03", "role": "main"},
{"chip_id": "board_04", "role": "readout"},
])
exp.set_calibration(cal)
rec = exp.new_recording("main", dt=0.001)
exp.log_metrics({"rate": 18.4, "selectivity": 0.87})
exp.save("./experiments") # JSON + HDF5Visualization
Spike rasters, membrane voltage traces, firing rate distributions, population activity, synaptic weight matrices, and multi-experiment comparisons. One-line dashboard for any recording. All plots composable with matplotlib.
import nuro.plot as nplot
nplot.spike_raster(spikes, dt=0.001)
nplot.voltage_traces(voltages, dt=0.001, neuron_indices=[0, 1, 2])
nplot.firing_rates(spikes, duration=0.5, dt=0.001)
nplot.weight_matrix(weights)
nplot.experiment_dashboard(rec, save_path="dashboard.png")
nplot.compare_recordings({"chip_A": rec_a, "chip_B": rec_b})Hardware Adapters
Bridge any data source into Nuro recordings. GPU training results, Loihi monitors, SynSense event streams, CSV files, HDF5, numpy arrays, NIR files, and AER binary formats. No vendor lock-in. Offline analysis without hardware SDKs.
from nuro.adapters.samna import SamnaAdapter
from nuro.adapters.aer import from_aedat
rec = SamnaAdapter.from_events(events, num_neurons=128, dt=1e-3)
rec = from_aedat("recording.aedat", num_neurons=256, dt=1e-3)
rec = nuro.adapters.from_csv("data.csv", dt=1e-3)
rec = nuro.adapters.from_numpy(spike_array, probe_name="spikes")Get Started
# DevTools only (numpy + h5py + matplotlib)
pip install nuro[devtools]
# Full stack (DevTools + GPU training + hardware deployment)
pip install nuro[gpu,devtools]
# Quick test
python -c "
import numpy as np
from nuro.adapters.aer import from_aer_events
from nuro.calibration import CalibrationProfile
from nuro.sweep import ParameterSweep
import nuro.plot as nplot
# Import events, calibrate, sweep, plot
rec = from_aer_events(np.array([0,1,0,2]), np.array([.001,.002,.005,.009]), num_neurons=3, dt=1e-3)
nplot.spike_raster(rec.get('spikes'), dt=1e-3)
print('Nuro DevTools working.')
"Stop losing experiment data in ad-hoc scripts.
Open source. Apache 2.0. Works with any chip that produces spike events. Cloud experiment storage coming in v0.9.